cellNexus is a query interface for programmatic exploration and retrieval of harmonised, curated, and reannotated CELLxGENE human-cell-atlas data.
This standalone documentation website provides:
- Detailed data-processing information (quality control, harmonisation, and expression representations).
- Complete explanations of metadata columns used in filtering and interpretation.
- Guided examples for metadata-first exploration and gene-expression analysis.
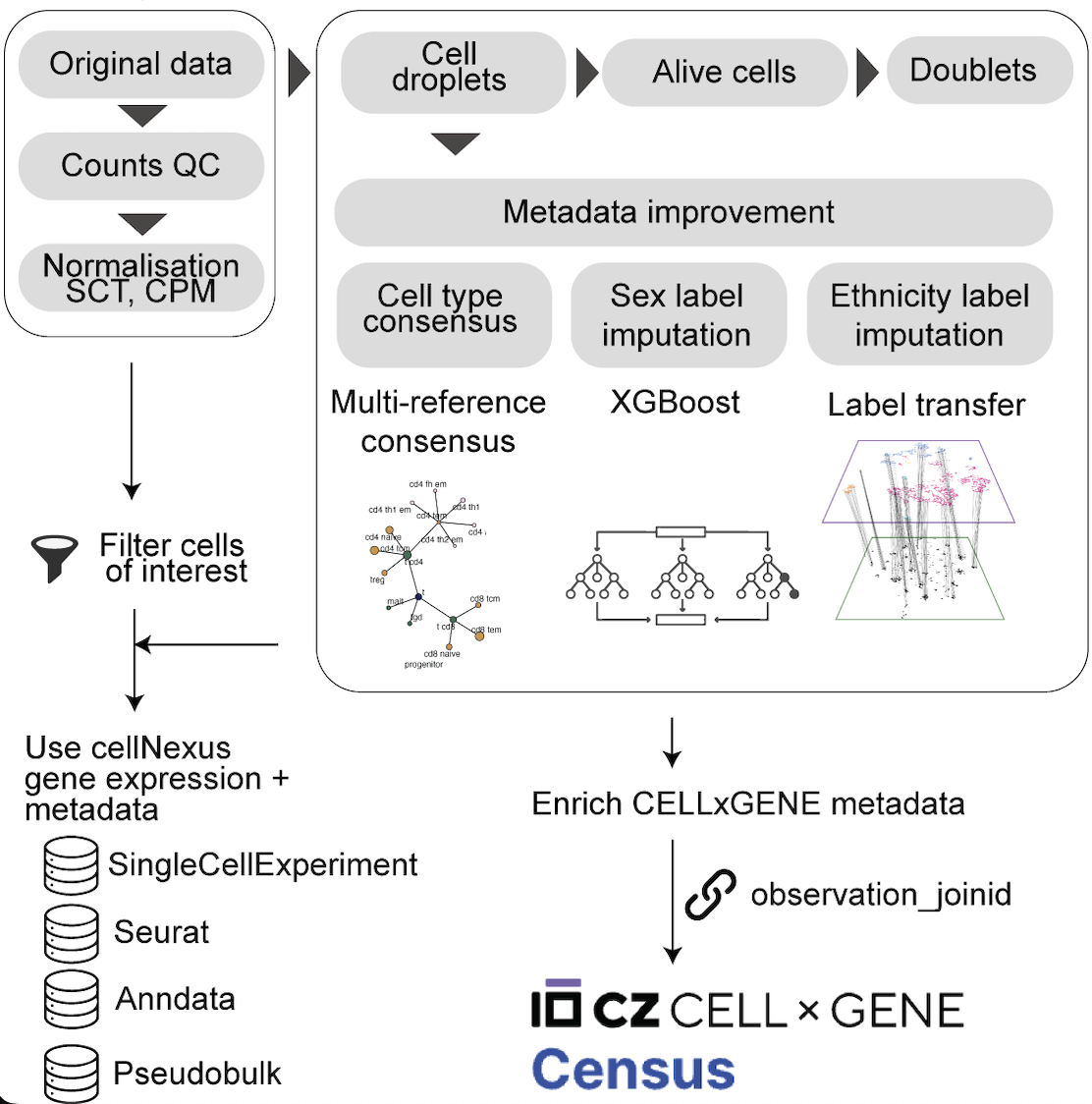
Data Processing Overview
The harmonisation pipeline standardises data across datasets so queries are consistent across studies:
- Metadata are retrieved from cloud-hosted harmonised tables.
- Standardised quality control removes empty droplets, dead/damaged cells, and likely doublets.
- Cell-level data are served through common assay layers (
counts,cpm,sct,pseudobulk). - Outputs are returned in analysis-ready R formats such as
SingleCellExperimentandSeurat.
Quality control steps
The QC flags used throughout cellNexus are computed using HPCell. In brief: - Empty droplets (empty_droplet): - Computed from a SingleCellExperiment. - Excludes mitochondrial genes and ribosomal genes before scoring: - Computes, per cell, the number of expressed genes and flags a cell as an empty droplet when (n_ <) RNA_feature_threshold (by default 200, except for targeted panels such as Rhapsody technology).
- Alive cells (
alive):- Filters out empty droplets.
- Computes per-cell QC metrics from raw counts using
scuttle::perCellQCMetrics(..., subsets=list(Mito=...)), where the mitochondrial subset is defined by^MTin the feature names. - Determines high mitochondrial content via
scater::isOutlier(subsets_Mito_percent, type="higher"). Outlier calling is performed within each cell-type group using our harmonised labelcell_type_unified_ensemble. - Alive cells are labelled as those without high mitochondrial content (
!high_mitochondrion).
- Doublets (
scDblFinder.class):- Filters out empty droplets.
-
scDblFinder::scDblFinder()default parameters are used. For cells that cannot be classified byscDblFinder, the class is set to"Unknown"to avoid dropping cells.
RNA abundance
- RNA counts:
- RNA count distributions per sample are annotated from cellxgenedp, using the
x_approximate_distributioncolumn.
- RNA count distributions per sample are annotated from cellxgenedp, using the
- CPM:
- Counts-per-million normalisation computed from the raw counts assay via
scuttle::calculateCPM().
- Counts-per-million normalisation computed from the raw counts assay via
- Rank:
- Per-cell gene-expression ranks computed with
singscore::rankGenes(). - Implemented in column chunks (default 1000 cells per slice) to handle very large datasets; slices are written to disk as an
HDF5Array-backed sparse integer matrix and then column-bound.
- Per-cell gene-expression ranks computed with
- SCT:
- Variance-stabilising normalisation computed with Seurat
SCTransform()(v2), with regression of cell-level covariates (subsets_Mito_percentandsubsets_Ribo_percent). - QC filtering is applied first.
- The median common scale across the whole resource is applied (scale_factor=2186)
- Variance-stabilising normalisation computed with Seurat
- Pseudobulk:
- All low-quality cells flagged by QC are removed before aggregation.
- Aggregates counts across cells using
scuttle::aggregateAcrossCells(), aggregatingsample_idand the harmonised cell type (cell_type_harmonised_ensemble).
Metadata Explore
Detailed field definitions for the CELLxGENE schema follow the CELLxGENE schema 5.1.0, and CELLxGENE Census schema
Dataset-level columns
| Column | Description |
|---|---|
dataset_id |
Primary dataset identifier in the atlas. |
collection_id |
Parent collection grouping related datasets. |
citation |
Citation or reference for the dataset. |
dataset_version_id |
Versioned identifier for a dataset release. |
explorer_url |
URL to browse the dataset in CELLxGENE explorer. |
filesize |
Reported size of source data files. |
filetype |
Storage format of source expression files. |
published_at |
Publication timestamp for dataset release. |
raw_data_location |
Location of raw data upstream. |
revised_at |
Last revision timestamp for dataset metadata. |
schema_version |
CELLxGENE schema version for this record. |
suspension_type |
Suspension type (e.g. cell vs nucleus). |
title |
Dataset publication title. |
url |
Dataset H5AD URL. |
donor_id |
Donor identifier. |
Sample-level columns
| Column | Description |
|---|---|
assay |
Sequencing technology label. |
assay_ontology_term_id |
Ontology identifier for assay. |
development_stage |
Developmental stage label. |
development_stage_ontology_term_id |
Ontology identifier for development_stage. |
self_reported_ethnicity |
Self-reported ancestry or ethnicity label. |
self_reported_ethnicity_ontology_term_id |
Ontology identifier for self-reported ethnicity. |
experiment___ |
Upstream experiment grouping variable. |
organism |
Organism name (e.g. human). |
organism_ontology_term_id |
Ontology identifier for organism. |
sex |
Recorded biological sex label. |
sex_ontology_term_id |
Ontology identifier for sex. |
tissue |
Tissue label. |
tissue_type |
Tissue class (e.g. tissue vs organoid). |
tissue_ontology_term_id |
Ontology identifier for tissue. |
disease |
Disease annotation. |
disease_ontology_term_id |
Ontology identifier for disease. |
is_primary_data |
Whether observations are marked as primary (avoid duplicated cells). |
Cell-level columns
| Column | Description |
|---|---|
cell_id |
Cell identifier. |
cell_type |
Cell-type label from CELLxGENE Census API. |
cell_type_ontology_term_id |
Ontology identifier for cell_type. |
observation_joinid |
Cell ID join key linking metadata. |
Harmonised and custom columns (cellNexus)
Through harmonisation and curation, cellNexus adds columns that are not present in the original CELLxGENE metadata alone.
| Column | Description |
|---|---|
cell_count |
Number of cells in a dataset. |
feature_count |
Number of genes in a dataset. |
age_days |
Donor age in days. |
tissue_groups |
Coarse tissue grouping for analysis. |
empty_droplet |
Quality-control flag for empty droplets. |
alive |
Quality-control flag for viable cells (e.g. mitochondrial signal). |
scDblFinder.class |
Quality-control flag for doublet classification from scDblFinder. |
cell_type_unified_ensemble |
Consensus immune identity from Azimuth and SingleR (Blueprint, Monaco). |
cell_annotation_azimuth_l2 |
Azimuth cell annotation. |
cell_annotation_blueprint_singler |
SingleR annotation (Blueprint). |
cell_annotation_blueprint_monaco |
SingleR annotation (Monaco). |
sample_heuristic |
Internal sample subdivision helper. |
file_id_cellNexus_single_cell |
Internal file id for single-cell layers. |
file_id_cellNexus_pseudobulk |
Internal file id for pseudobulk layers. |
sample_id |
Harmonised sample identifier. |
nCount_RNA |
Total RNA counts per cell (sample-aware). |
nFeature_expressed_in_sample |
Number of expressed features per cell. |
ethnicity_flagging_score |
Supporting score for ethnicity imputation. |
low_confidence_ethnicity |
Supporting flag for low-confidence ethnicity calls. |
.aggregated_cells |
Post-QC cells combined into each pseudobulk sample. |
imputed_ethnicity |
Imputed ethnicity label. |
atlas_id |
cellNexus atlas release identifier (internal use). |
is_immune |
Curated flag for immune-cell context. |
Client Usage Examples
R client (cellNexus)
library(cellNexus)
library(dplyr)
library(stringr)
metadata <- get_metadata() |>
join_census_table()
metadata <- metadata |>
keep_quality_cells()
query <- metadata |>
filter(
self_reported_ethnicity == "African",
str_like(assay, "%10x%"),
tissue == "lung parenchyma",
str_like(cell_type, "%CD4%")
)
sce <- get_single_cell_experiment(query, assays = c("counts", "cpm"))
pb <- get_pseudobulk(query)Python client (cellNexusPy)
Python support is available in the companion repository: MangiolaLaboratory/cellNexusPy.
from cellnexuspy import get_metadata, get_anndata
sample_dataset = "https://object-store.rc.nectar.org.au/v1/AUTH_06d6e008e3e642da99d806ba3ea629c5/cellNexus-metadata/sample_metadata.1.3.0.parquet"
conn, table = get_metadata(parquet_url=sample_dataset)
table = table.filter("""
empty_droplet = 'false'
AND alive = 'true'
AND "scDblFinder.class" != 'doublet'
AND feature_count >= 5000
""")
query = table.filter("""
self_reported_ethnicity = 'African'
AND assay LIKE '%10%'
AND tissue = 'lung parenchyma'
AND cell_type LIKE '%CD4%'
""")
adata = get_anndata(query, assay="cpm")
pb = get_anndata(query, aggregation="pseudobulk")
conn.close()For other implementation details and code examples, see cellNexus README
